Self-Organizing Maps (SOM)

Hey folks, in another data science post today, we will learn about Self-Organizing Maps (SOMs). Before you proceed to have a thorough grip on ANN’s and Clustering algorithms and also read Wikipedia’s vector quantization. Kohonen proposed this technique, therefore, it is called a Kohonen map. It is based on biological neural networks.
What is Self-Organizing Map?
SOM is a powerful artificial neural network that operates on the principle of unsupervised learning. Unsupervised learning does not have labeled data which helps an algorithm identify something about the data like a class label. But, we do not have such a thing as labeling the data here. SOM comes in handy here. Scientifically, SOM is used in many fields like signal processing, identifying patterns, detecting earthquakes(a real, lifesaver). A SOM takes a high dimensional dataset and reduces it to a low-level dimension. Kohonen map is neither a feed-forward nor a feed-back it is a self-organizing neural network.
How Does SOM work?
First, let’s understand K-means.
- Take a datapoint from infinite data points.
- Identify the closest data points to it by any means like mean distance, max distance, centroid
- Keep clustering until you cover maximum data points.
Self-organizing maps apply competitive learning.
Learning in SOM:
From here on, the post will get a little complex. Bear with me I’ll try to explain in a simple manner all you have to do is get the basics right. As we humans respond to environmental stimuli like cold, heat, air the SOM also responds to the input data. Let’s say our input data is some earthquake activity dataset. The SOM will take many factors into consideration like sea level, tectonic plates, faults, etc. It analyzes different countries’ different landscape and let’s say plate tectonics are the most important features and it tends to compress towards them. This does not happen immediately, it happens after a long cycle where there is competition for other features where there is a maximum focus in each cycle.
The above process is called self-organizing.
Training in Self-Organizing Maps
Dataset is fed to the network, its Euclidean distance to all weight vectors is computed.
Best matching unit (BMU) is a neuron which has a weight vector the most similar to the input.
These weights of the BMU and neurons close to it in the SOM grid are adjusted towards the input vector. We change the magnitude with time and the grid-distance from the BMU.
Neighborhood function Θ(u, v, s) depends on the grid-distance between the BMU(u) and a neuron v. Let’s say, it is 1 for all neurons close enough to BMU and for others, but we prefer a Gaussian function as a practice. Regardless of the functional form, the neighborhood function shrinks with time. When the neighborhood is broad, the self-organizing takes place on a large scale. After it gradually shrinks to just a couple of neurons, then weights converge to local estimates. Sometimes, the learning coefficient α and the neighborhood function Θ decrease steadily with increasing s, in others, they decrease in a gradual fashion, once every T steps.
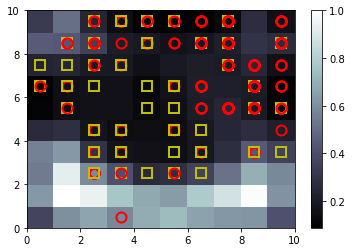
Self Organizing Maps Algorithm
- Randomize the node weight vectors in a map
- Randomly pick an input vector
- Traverse each node in the map
- Use the Euclidean distance formula to find the similarity between the input vector and the map’s node’s weight vector
- Track the node that produces the smallest distance (this node is the best matching unit, BMU)
- Update the weight vectors of the nodes in the neighborhood of the BMU (including the BMU itself) by pulling them closer to the input vector
- Increase s and repeat from step 2.
Thank you, in the next post I will do a programming example of this algorithm.
Leave a Reply